When we hear about bacteria, it is usually linked to disease: an Escherichia coli outbreak tied to romaine lettuce and the rise of antibiotic-resistant bacterial strains are just two examples. However, bacteria are also connected with valuable properties like lower risk of inflammatory bowel disease, diabetes, obesity, and other chronic diseases. Most animals have a mutual symbiotic relationship, meaning that both organisms benefit from their association, with at least a few bacterial species.
Our colonized microorganisms are collectively referred to as the “microbiota.” While scientists have known for decades that we are host to millions of bacteria, research investigating the role of the microbiome has only recently become a focal point. It has expanded to encompass not only human medicine but the health of other animals as well (including pets), leading to astounding links to our overall health.
Origins of Microbiome Research
The idea of the microbiome first arose when scientists discovered that removing all microorganisms from animals, called germ-free1,2, created individuals that were physiologically different than their counterparts who played host to various bacterial species. For example, germ-free mice require vitamin K supplements3, as they cannot metabolize it from their food. They also tend to be more susceptible to infection by some bacteria4 and viruses5.
Thus, it seemed that microbes were important regulators of animal nutrition, health, and physiology. Researchers called this collection of microscopic organisms the “normal flora” or “microflora.” These terms have since fallen out of favor, as most current researchers refer to our collective bacterial symbionts as the “microbiota” or the “microbiome.”
Human Microbiome Research
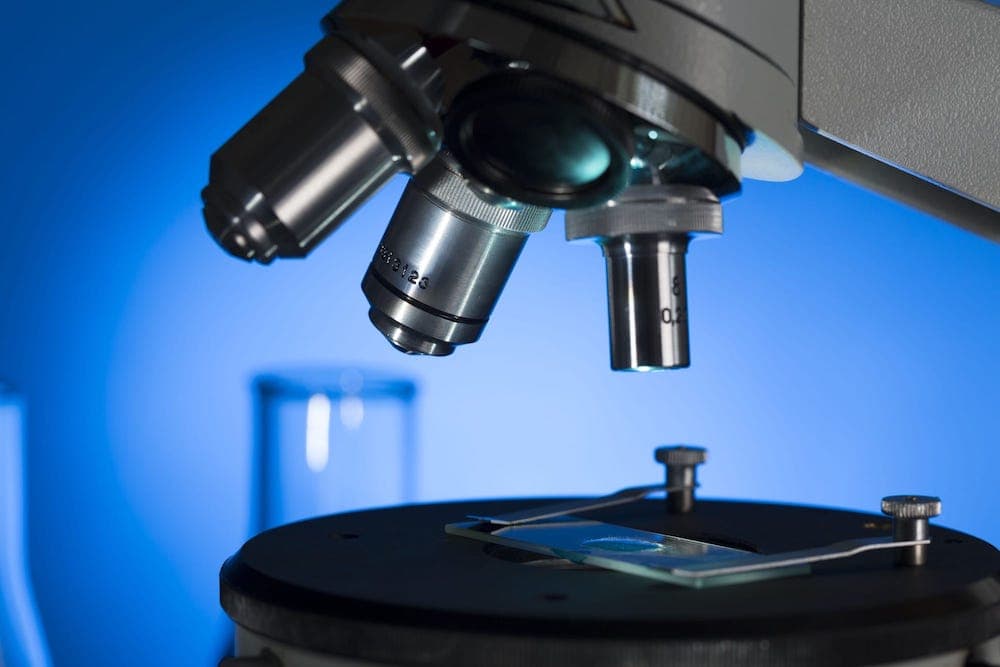
For a long time, animal microbiomes, including that of humans, remained poorly understood as researchers were limited by available technologies. Culture-based methods were the most popular way to determine which bacterial symbionts thrived in different body cavities. “Culture-based” means that researchers grow bacteria taken from a specific microbiome, such as a mouth swab, and attempt to grow them in the lab and identify them. They do this by providing the necessary components for growth, such as nutrient-rich liquid or solid agar plates.
There are a variety of methods by which these bacteria can be identified, for example, the bacteria may be able to utilize a certain resource, resulting in a color change in the broth. It is a way to identify specific bacterial species within a sample, much like identifying a toothpick in a sea of needles by determining which one catches on fire. The main limitation of this method is that the vast majority of bacteria simply cannot be grown in the lab, thus limiting the number of species that can be identified.
The success of the Human Genome Project6 in 2003 transformed our ability to classify microbial companions in part by advancing DNA sequencing technologies, which are non-culture-based, and therefore not subject to the limitations of culturing.
DNA sequencing, the practice of determining the genetic composition of an organism’s DNA (our hereditary material), is much like reading a book. DNA sequences are written in a code of 4 chemical bases which, because they are unique between organisms, can help scientists ascertain the species. DNA is isolated by chemically breaking a cell open and purifying the DNA from the rest of the cell’s contents.
Microbial DNA can further be distinguished from host DNA (such as human) by sequencing a gene that the host does not have, called 16S ribosomal RNA (rRNA). Sequencing has been used for decades in scientific research but had not been efficient enough to sequence the astounding numbers of bacterial symbionts present in one individual (generally thought to be around 1012 cells7). The Human Genome Project, which revealed an entire human DNA sequence, demonstrated that such a feat was achievable, thus leading to the initiation of the Human Microbiome Project from 2008 - 2013.
The goal of this project was to create a database describing all the microorganisms living in association with humans. They found that the human microbiome is incredibly diverse between healthy people, likely influenced by diet, environment, genetics, and exposure. These findings led the way for more research to characterize the functional consequences of our microbiome.
Numerous studies, performed both before and after the Human Microbiome Project, have described the impacts of our varied microbial populations on health. In 2018 alone, over 1,000 microbiome related studies have been published. Our microbiome affects a myriad of physiological conditions. Oral bacteria, for instance, can protect teeth from cavities by outcompeting the species Streptococcus mutans9.
In the gut microbiota, bacteria aid in digestion of nutrients10 but can also influence risk factors for numerous disease states11 such as obesity, inflammatory bowel disease, or even allergies12. Recent work has also found preliminary ties between gut microbes and psychiatric disorders13, such as depression and anxiety, as well as circadian rhythms14, our sleep/wake cycles. The more scientists uncover about our symbiosis with microorganisms, it becomes clearer that the health of most life on Earth is dependent to some extent on bacteria.
Research In Other Animals
Knowledge of the human microbiome is generally supplemented by research done on other animals. Mice15, for instance, are commonly used in microbiome research, especially as germ-free animals. However, we are also learning more about the microbiomes of other animals, specifically as researchers broaden the scope of these types of studies.
For example, the microbiomes of dolphins at the Shedd Aquarium in Chicago16 are affected more by diet and the air they breathe than water composition in their tank or skin-to-skin contact with their handlers, and captive baboons have vastly different microbiomes than wild baboons17. Understanding the differences in composition between wild animals and those in captivity can help conservation efforts to keep captive animals healthy even when introduced to spaces that could potentially alter their microbiota.
Microbiome research has additionally explored differences between our close canine companions and their relatives, wolves18. Domestic dogs and wolves do share some bacterial groups, but a number of bacteria are specific to each species. The significance of these differences has not yet been explored, but it suggests that the domestication of dogs may have impacted their microbial communities, as well as their ability to tolerate small amounts of starch and vegetation in their diets19.
Dog Pet microbiome research has slowly been advancing and the rate of increase in previous years has been substantial. Previous work has focused on how diet shapes your dog’s gut bacteria and the impact of gut microbes on nutrition. However, there are broad efforts to determine how pet parents can keep their dogs healthy. Feline microbiomes are also under scrutiny20,21, although to a lesser extent than dogs. The bacteria that comprise their gut have been identified and are similar to other mammals. Moreover, this composition can be influenced by their diet.
The Future of Microbiome Research
The more we know about the world at large, including animals kept in captivity or our companion animals, the better we can keep them (and ourselves) healthy and ensure they receive optimal care. Microbiome studies have come a long way over the past two decades. Advances in sequencing technologies have allowed scientists to conduct research that is more comprehensive than in the past.
However, correlation is not causation and most relationships between health or disease risk and microbial composition has been associative, though databases are increasing rapidly due to the influx of current research. Further technological advances will provide direct links between specific bacterial species and host health. In-depth knowledge of these relationships will also lead to the development of improved diagnostic and therapeutic tools to diagnose and treat diseases caused by an unbalanced microbiota.
References
1. Luckey, T. D. Effects of Microbes on Germfree Animals11Presented in modified form as the principal talk at the International Symposium on Microecology, Berlin, September, 1964. in Advances in Applied Microbiology (ed. Umbreit, W. W.) 7, 169–223 (Academic Press, 1965).
2. Al-Asmakh, M. & Zadjali, F. Use of Germ-Free Animal Models in Microbiota-Related Research. J. Microbiol. Biotechnol. 25, 1583–1588 (2015).
3. Hirayama, K., Uetsuka, K., Kuwabara, Y., Tamura, M. & Itoh, K. Vitamin K deficiency of germfree mice caused by feeding standard purified diet sterilized by gamma-irradiation. Exp. Anim. 56, 273–278 (2007).
4. Khosravi, A. et al. Gut microbiota promote hematopoiesis to control bacterial infection. Cell Host Microbe 15, 374–381 (2014).
5. Pfeiffer, J. K. & Sonnenburg, J. L. The intestinal microbiota and viral susceptibility. Front. Microbiol. 2, 92 (2011).
6. An Overview of the Human Genome Project. National Human Genome Research Institute (NHGRI) Available at: https://www.genome.gov/about-genomics/educational-resources/fact-sheets/human-genome-project. (Accessed: 5th December 2018)
7. Sender, R., Fuchs, S. & Milo, R. Are We Really Vastly Outnumbered? Revisiting the Ratio of Bacterial to Host Cells in Humans. Cell 164, 337–340 (2016).
8. NIH Human Microbiome Project - About the Human Microbiome. Available at: https://hmpdacc.org/hmp/overview/. (Accessed: 5th December 2018)
9. Huang, X. et al. A Highly Arginolytic Streptococcus Species That Potently Antagonizes Streptococcus mutans. Appl. Environ. Microbiol. 82, 2187–2201 (2016).
10. Flint, H. J., Scott, K. P., Louis, P. & Duncan, S. H. The role of the gut microbiota in nutrition and health. Nat. Rev. Gastroenterol. Hepatol. 9, 577–589 (2012).
11. Clemente, J. C., Ursell, L. K., Parfrey, L. W. & Knight, R. The impact of the gut microbiota on human health: an integrative view. Cell 148, 1258–1270 (2012).
12. Huffnagle, G. B. The microbiota and allergies/asthma. PLoS Pathog. 6, e1000549 (2010).
13. Evrensel, A. & Ceylan, M. E. The Gut-Brain Axis: The Missing Link in Depression. Clin. Psychopharmacol. Neurosci. 13, 239–244 (2015).
14. Thaiss, C. A. et al. Microbiota Diurnal Rhythmicity Programs Host Transcriptome Oscillations. Cell 167, 1495–1510.e12 (2016).
15. Clavel, T., Lagkouvardos, I., Blaut, M. & Stecher, B. The mouse gut microbiome revisited: From complex diversity to model ecosystems. Int. J. Med. Microbiol. 306, 316–327 (2016).
16. Researchers study how environment affects dolphin microbiomes at Shedd Aquarium. Available at: https://phys.org/news/2018-07-environment-affects-dolphin-microbiomes-shedd.html. (Accessed: 5th December 2018)
17. Baboons shed light on antimicrobial resistance. Available at: https://phys.org/news/2018-06-baboons-antimicrobial-resistance.html. (Accessed: 5th December 2018)
18. Wu, X. et al. Analysis and comparison of the wolf microbiome under different environmental factors using three different data of Next Generation Sequencing. Sci. Rep. 7, 11332 (2017).
19. Moon, C. D., Young, W., Maclean, P. H., Cookson, A. L. & Bermingham, E. N. Metagenomic insights into the roles of Proteobacteria in the gastrointestinal microbiomes of healthy dogs and cats. Microbiologyopen e00677 (2018).
20. Kitty Biome. Available at: https://www.kittybiome.com/. (Accessed: 5th December 2018)
21. Minamoto, Y., Hooda, S., Swanson, K. S. & Suchodolski, J. S. Feline gastrointestinal microbiota. Anim. Health Res. Rev. 13, 64–77 (2012).